Can We Trust The Results From Popular DNA Tests?

Contents
DNA testing errors happen
So to do that we’re going to step back a little and look at basic science, and how it applies to genetic testing. In an ideal world you’d provide your sample send it off to the lab where it would be tested giving you your results which you could be sure were 100% accurate. Sadly, that will never happen, there’s no such thing as a perfect test. Simple technical issues can have profound effects, for example what if the composition of the testing solution is slightly different, the pH is changed, the room is 5 degrees colder… all this can impact outcomes. We try to minimize this by controlling for as many things as possible but it’s not a perfect science.False positives in DNA testing
These errors, and imperfections that are present in the test itself give rise to errors in reporting. For example, you may be CC (no “mutation”) for the T47C SNP in SOD2 but a test reports you as CT. At this point, we’re getting into the realm of false positives and false negatives, and understanding these and how to control for them is useful for understanding wider science. In the above example, we know that “T” is the risk allele. You are actually CC but have been reported CT, this is an example of a false positive. Basically, you’re being made to worry about a mutation you don’t have.False negatives in DNA testing
Flipping it around, let’s say you were actually CT, but the test reported you as CC. This is a false negative, you think you’re in the clear but in effect you’re not. You have a “mutation” that isn’t being reported.Sensitivity and specificity of DNA testing
With this information, and information about the population, we can determine what the sensitivity and specificity of a test are. Sensitivity is defined as the number of true positives/false negatives. Sensitivity therefore is a measure of how accurately we pick up people who are actually positive. Specificity by contrast is a measure of true negatives/false positives. Specificity is therefore a measure of how accurately we pick up people who are actually negative. So, a test with perfect specificity and selectivity would correctly assign people 100% of the time. As I said above, this test doesn’t exist… but there are some things we can do to mitigate it. The first being repetition.Repetition and aggregation
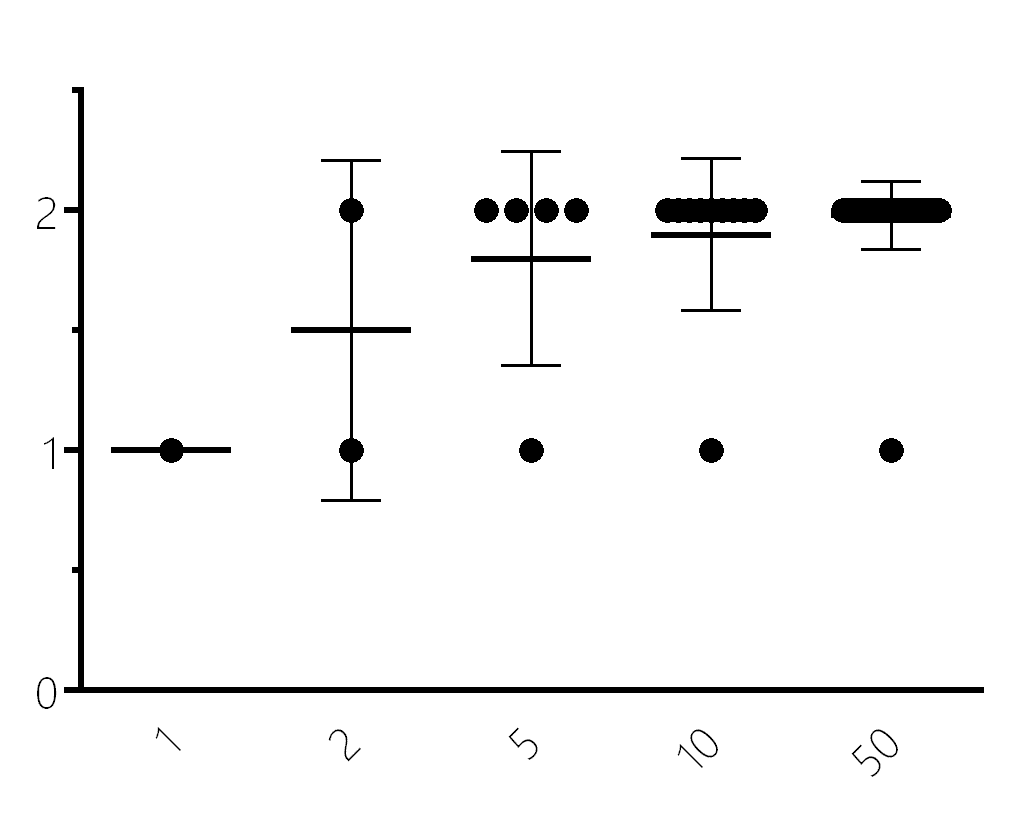
More consumer genetic tests = more accuracy
You can think about it like this: if you have 23andme data, then utilize one of our testing kits at Gene Food, where both SNPs match you can be more sure of the result than from just one test. Of course the question is then, what if I get two different results? Three tests… five tests? The cost goes up and the likelihood of an individual using the test goes down. Repetition and aggregation are expensive, and as tests go, genotyping is actually very accurate for most common polymorphisms, but…DNA testing accuracy rates
But how does that accuracy stack up against the big number of SNPs we get back in our reports? This paper 2 suggests that for common SNPs an accuracy of over 99% can be assumed. Let’s be generous and call that 99.99% to be generous. Where basically 1 in every 10,000 SNPs would be wrong. Your average consumer genetics kit reports anywhere between 500,000 and 1,000,000 SNPs so if we apply this number we’re looking at approximately 50-100 incorrect SNPs, and that was me being generous with the percentages, if we shift it back to the 99% stated we get 5-10,000 incorrect SNPs!Rare alleles are over-reported
The paper also points out that rare alleles, i.e. “mutations,” or SNPs, where the majority of people will have one allele, but a very small population will have another are even more susceptible to error. In the paper I reference above, they talk about a significant over prediction of positive rare alleles which were not confirmed with a more robust sequencing approach. Maybe not the end of the world if you’re using your report to investigate your ancestry, or make dietary changes, but as we know people will suggest huge lifestyle and dietary changes on just a single SNP! If you want to get an idea of the frequency of a SNP we report the allele frequency on all of our gene pages.Are consumer genotyping services worthless?
So I bet you’re wondering, why is the guy who writes on a website advocating the use of consumer genetic testing to inform nutrition and DNA diets being so down on the state of the industry? Well, if you listen to the podcast, you’ll know that I’m a perennial fence sitter. But in reality, I’m not being down. We recognize the weaknesses inherent in the tests, and we’ve tried to incorporate that into our ethos. So you won’t find us pushing extreme diets based on a single SNP that you carry. Firstly, because we don’t really push an extreme diet at either end of our nutrition plan. Secondly because we would rather take the approach of weighting SNPs, then grouping them together and using this to make a conclusion based on a comprehensive score, instead of based on one marker. We also performed a thorough investigation into genotyping chip suppliers and picked the one that provided both the best coverage, but also the best quality of coverage for our needs. We’re not saying our test is better, just that it’s better for us. Our lab uses a Screening Array manufactured by Illumina, which is pretty much the best out there.Summary
- Take multiple tests… not a serious option but if you’re worried about something at a clinical level this is what you will do with your healthcare provider.
- Consider investing in a newer consumer technologies such as whole exome or genome sequencing, which should feature fewer errors.
- Don’t make significant changes based on a single report of a single SNP.
- Look at your SNPs in the context of other SNPs. One may be wrong, but if you assess 10 to make a lifestyle change your chances of it being wrong are significantly lower.