The Science Behind DNA Diets
A growing body of scientific research is confirming what many of us have always known intuitively. There is no “one size fits all” when it comes to diet.
While DNA has a large role to play in personalizing nutrition, it is not the only factor. The best personalized nutrition regimens tailor diet, lifestyle, and exercise to genetic predispositions, and that is what we do at Gene Food. For example, we know that 50% of the post-meal blood sugar response is genetically determined, however, sleep and exercise also play a role. By pairing nutrition insights with sleep chronotype and exercise tips, we offer a comprehensive approach. Our job is to highlight the most important genetic markers that impact on food choices so our customers can better understand why one diet may work for them, while another likely won’t. In order to do this, we evaluate hundreds of peer reviewed studies and evaluate the research using a scoring system we call Science Grade. Genes with the most scientific validation receive the greatest weight in our algorithm, whereas genes with less research receive less weight. Our research team utilizes an inclusive research process which ensures that the health impact for underrepresented communities, the elderly, women, and people of color are all factored into the research behind our products and our content.
Our Scoring System
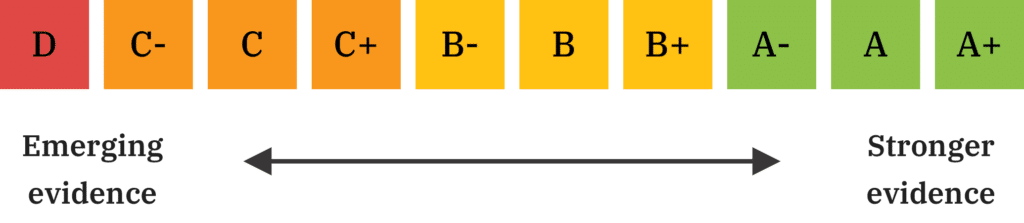
In Defense of DNA diets
Have you been Googling “do DNA diets work?”
There is new and exciting research from peer reviewed journals on the subject of nutrigenomics every month. Read our summary of all the reasons tailoring diet to DNA has benefit and will continue to evolve as research progresses.

Gene Food utilizes polygenic risk scoring
There is not one uniform DNA diet or one gene that determines health outcomes, and yet many of the studies billed as “DNA diet studies” look only at a single gene. At Gene Food, we use a comprehensive approach, called polygenic risk scoring, to assign diet types based on multiple genes and studies. Below, we list several of the most significant studies in our database (lasted updated 05/27/25).

Important Studies on DNA Diets
When evaluating the studies below several factors were considered. Repeatability of the studies, sample population sizes, assessment of outcomes in control individuals vs. variant-carrying individuals, test statistic values for associations between genetic pathways and phenotypes, interpretation of results, and quality of genotyping in the study methods were among the primary criteria in selecting these genes.
Alcohol Metabolism
| Study | Journal | Genes | Diet Issue | Findings |
|---|---|---|---|---|
| Polymorphisms in Alcohol Metabolism Genes ADH1B and ALDH2, Alcohol Consumption and Colorectal Cancer | PLOS One | ADH1B | Alcohol Metabolism and Colorectal Cancer | The SNP rs1229984 in the ADH1B gene had association with colorectal cancer risk. Those with this variant should be mindful of alcohol consumption, possibly eliminating it entirely from their diet (as it compounds CRC risk). |
Celiac Disease (Wheat)
| Study | Journal | Genes | Diet Issue | Findings |
|---|---|---|---|---|
| Association of LPP and TAGAP Polymorphisms with Celiac Disease Risk: A Meta-Analysis | International Journal of Environmental Research and Public Health | TAGAP | Celiac Disease (Wheat) | Across seven studies, there was a reported association between the TAGAP rs1738074 polymorphism and increased susceptibility to celiac disease; if this polymorphism is present in individuals, it is recommended to immediately switch to a gluten-free diet (along with other celiac disease management solutions). |
Heart Health
| Study | Journal | Genes | Diet Issue | Findings |
|---|---|---|---|---|
| Interaction of PPARG Pro12Ala with dietary fat influences plasma lipids in subjects at cardiometabolic risk | Journal of Lipid Research; published by the American Society for Biochemistry and Molecular Biology | PPARG | Cholesterol | PPARG-ProAla12 carriers tend to have higher mean total cholesterol and LDL-cholesterol concentrations on higher saturated fat diets than noncarriers. |
| APOA II genotypes frequency and their interaction with saturated fatty acids consumption on lipid profile of patients with type 2 diabetes | Clinical Nutrition | APOA2 | Lipid metabolism | With the APOA2 rs5058 ‘G’ allele variant indicative of poor lipid outcomes, especially in individuals at risk of developing Type 2 diabetes, those with the variant showed improved lipid outcomes comparable to individuals with the normal ‘A’ allele after being put on a diet with reduced saturated fat and carbohydrate consumption. |
| Apolipoprotein E phenotype regulates cholesterol absorption in healthy 13-month-old children–The STRIP Study | Pediatric Research | APOE | Cholesterol | Children with the apoE4 polymorphism have higher serum concentrations of diet-introduced plant sterols in early childhood than children who had the apoE 3/3 variant; a low-fat diet can reduce the risk of high serum cholesterol for individuals with the APOE4 polymorphism |
| Biomarkers of cholesterol homeostasis in a clinical laboratory database sample comprising 667,718 patients | Journal of Clinical Lipidology | APOE | Cholesterol | Among carriers with different APOE variants, those with the APOE ε4 allele were found to have higher prevalence of cholesterol hyperabsorption than individuals with other variants; this information can help physicians and nutritionists advise individuals with the higher-risk variant to switch to lipid-lowering diets. |
| Large-scale association analysis identifies new risk loci for coronary artery disease | Nature Genetics | ABCG8 | Cholesterol | At study-wide significance levels, the ‘T’ allele of T322+431C in the ABCG8 gene, was found to be one of several key genetic components in coronary artery disease. As this SNP contributes to increased LDL levels, individuals with this variant may benefit from a diet consisting of lower cholesterol levels and lipid content. |
| C667T and A1298C polymorphisms of methylenetetrahydrofolate reductase gene and susceptibility to myocardial infarction: A systematic review and meta-analysis | International Journal of Cardiology | MTHFR | Myocardial Infarction Risk | MTHFR A1298C polymorphisms were shown to be associated with myocardial infarction risk in African, North American, and elderly populations, while the C667T polymorphism was associated with a 63% increased risk of MI (vs. C667C) in an African population subgrouping, indicating that individuals with these SNPs should consider switching to low-fat diets with increased omega-3 fatty acid consumption to reduce risk of MI. C667T Polymorphism: There were a total of 47 case-control studies regarding the C667T polymorphism. Of these 47, there was a single study involving an African population. For the African population, it was shown that the “T” allele of the C667T polymorphism was associated with a 63% increased risk of MI compared to the “C” allele. The I2 (%) for the African subgroup, a measure that describes the percentage of variation across studies that is due to heterogeneity rather than chance, was less than 44% for all 5 models tested. |
| Dietary linoleic acid interacts with FADS1 genetic variability to modulate HDL-cholesterol and obesity-related traits | Clinical Nutrition | FADS1 | HDL cholesterol | In patients with the FADS1 rs174547 polymorphism, with higher linoleic acid intake there were lower HDL-cholesterol levels observed. Those with this minor allele variant may benefit from lower linoleic acid as part of their diets. |
| Periconception Red Blood Cell Folate and Offspring Congenital Heart Disease | Annals of Internal Medicine | MTHFR | Maternal folate levels | Higher maternal folate levels were associated with a lower risk for congenital heart defects in babies. Mendelian randomization was done using the methylenetetrahydrofolate reductase (MTHFR) C677T as the genetic instrument. |
Immune System
| Study | Journal | Genes | Diet Issue | Findings |
|---|---|---|---|---|
| Association of single nucleotide polymorphisms in the diamine oxidase gene with diamine oxidase serum activities | European Journal of Allergy and Clinical Immunology | AOC1 | Histamine Intolerance | The ‘T’ allele in the C47T SNP of the AOC1 gene has association with histamine intolerance and reduced di-amine oxidase activity; individuals that switched to a diet with minimal consumption of histamine-rich foods noticed improvement in their symptoms that had arisen from histamine intolerance. |
| Histamine and histamine intolerance | The American Journal of Clinical Nutrition | AOC1 | Histamine intolerance | DAO activity increases when there is administration of vitamin C and vitamin B6; diets rich in these vitamins can mitigate some of the symptoms that arise from histamine intolerance |
Lactose Intolerance
| Study | Journal | Genes | Diet Issue | Findings |
|---|---|---|---|---|
| Genetic predisposition for adult lactose intolerance and relation to diet, bone density, and bone fractures | Journal of Bone and Mineral Research | LCT | Lactose Intolerance | The LCT(T/C-13910) polymorphism was found to be associated with reduced milk calcium intake, and is an indicator of subjective milk intolerance. Those with this SNP will likely benefit from switching to a lactose-free diet. |
Metabolic and Weight Loss
| Study | Journal | Genes | Diet Issue | Findings |
|---|---|---|---|---|
| PREDICT-1: Human postprandial responses to food and potential for precision nutrition* | Nature Medicine | Multiple | Blood sugar | 50% of the post-meal blood sugar response is attributable to genetics. Post-meal triglycerides and insulin less impacted by genetic factors. |
| A comparison of a ketogenic diet with a LowGI/nutrigenetic diet over 6 months for weight loss and 18-month follow-up | BMC Nutrition | Multiple | Weight loss, cholesterol, blood sugar | A low GI nutrigenomic diet significantly outperformed a ketogenic diet for weight loss, cholesterol, HDL, and fasting blood glucose after an 18 month follow up period. |
| Enhanced long-term dietary change and adherence in a nutrigenomics-guided lifestyle intervention compared to a population-based (GLB/DPP) lifestyle intervention for weight management: results from the NOW randomised controlled trial | BMJ Nutrition, Prevention, and Health | Multiple | Weight loss, cholesterol, blood sugar | Nutrigenomics paired with traditional weight loss and diabetes management programs showed much greater long term adherence than one size fits all protocols. |
*PREDICT-1 Study gender/demographic makeup: Of the entire cohort of twins, 82% are female, and mean age is 59 (this could signify that age may play a factor in the study results). No information regarding the ethnic backgrounds of the participants in the PREDICT-1 study is provided.
