Genetics 101
Needing a refresher on genetics? We have you covered. In Genetics 101, we will cover everything you learned in high school to get you up to speed.
The code within your cells
Our body is made up of roughly 30 trillion cells, all with specialized functions, creating the diverse range of tissues and organs required to make a human. Within each cell, in a structure known as the nucleus, we have 23 pairs of chromosomes (hence the name of 23andme), which is 46 chromosomes in total. These 23 pairs of chromosomes are inherited from your parents, one from your mother and one from your father.
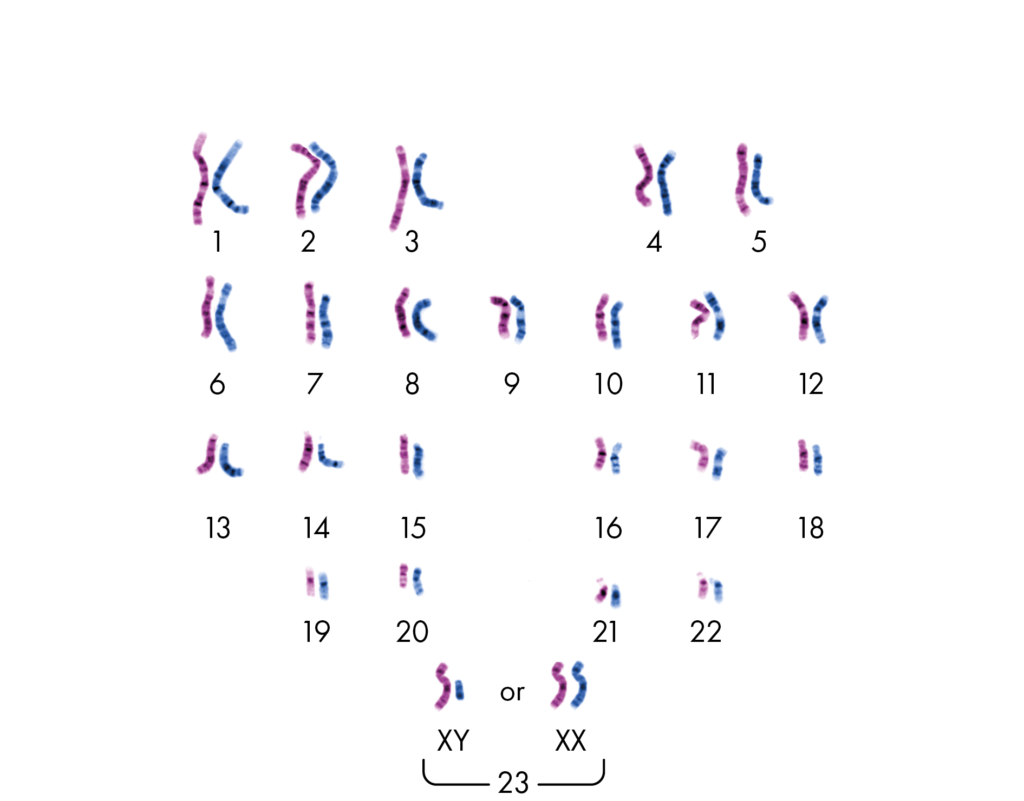
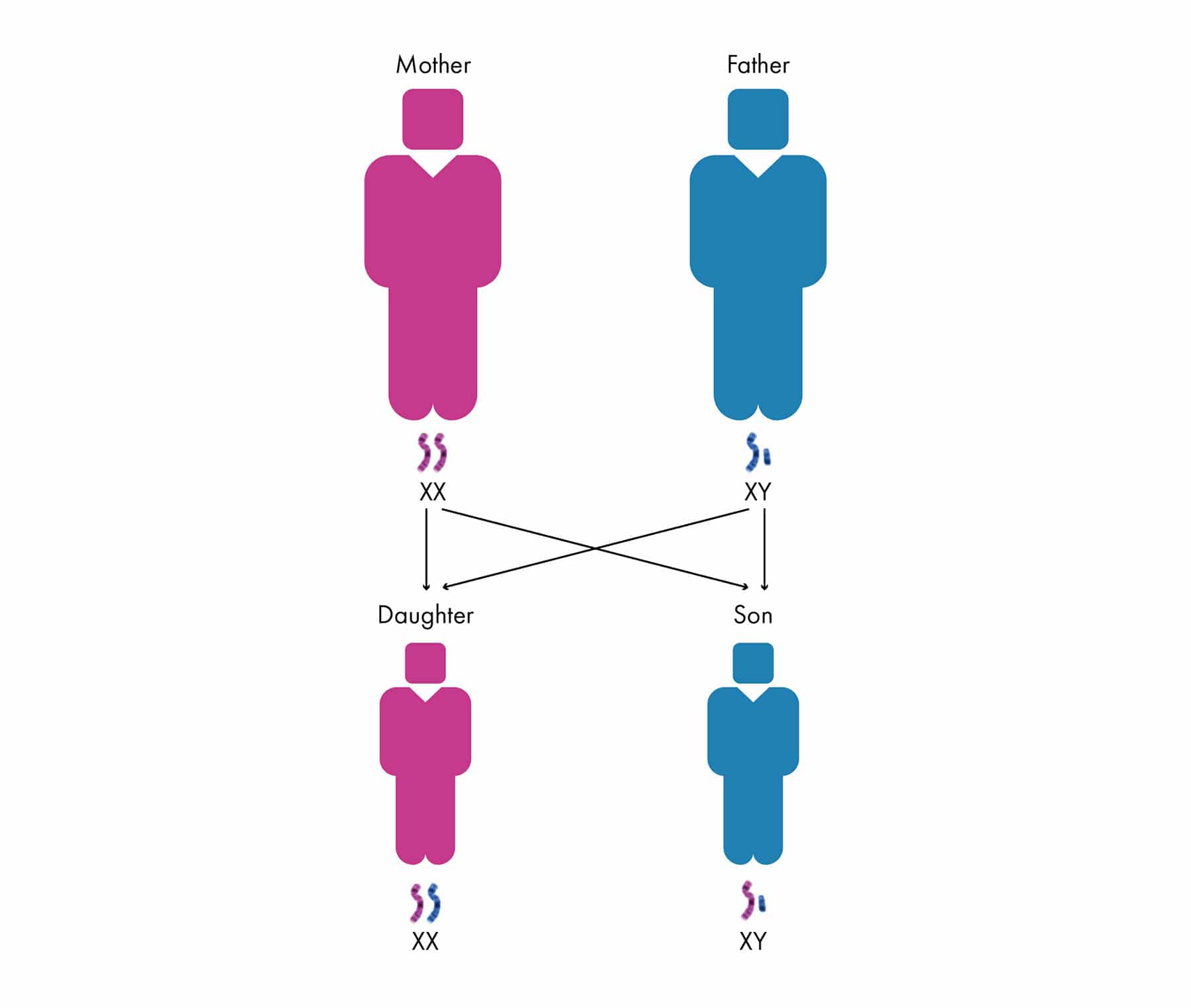
The sex chromosome pair determine whether you are male (XY) or female (XX). While the other 22 pairs of chromosomes are known as autosomes.
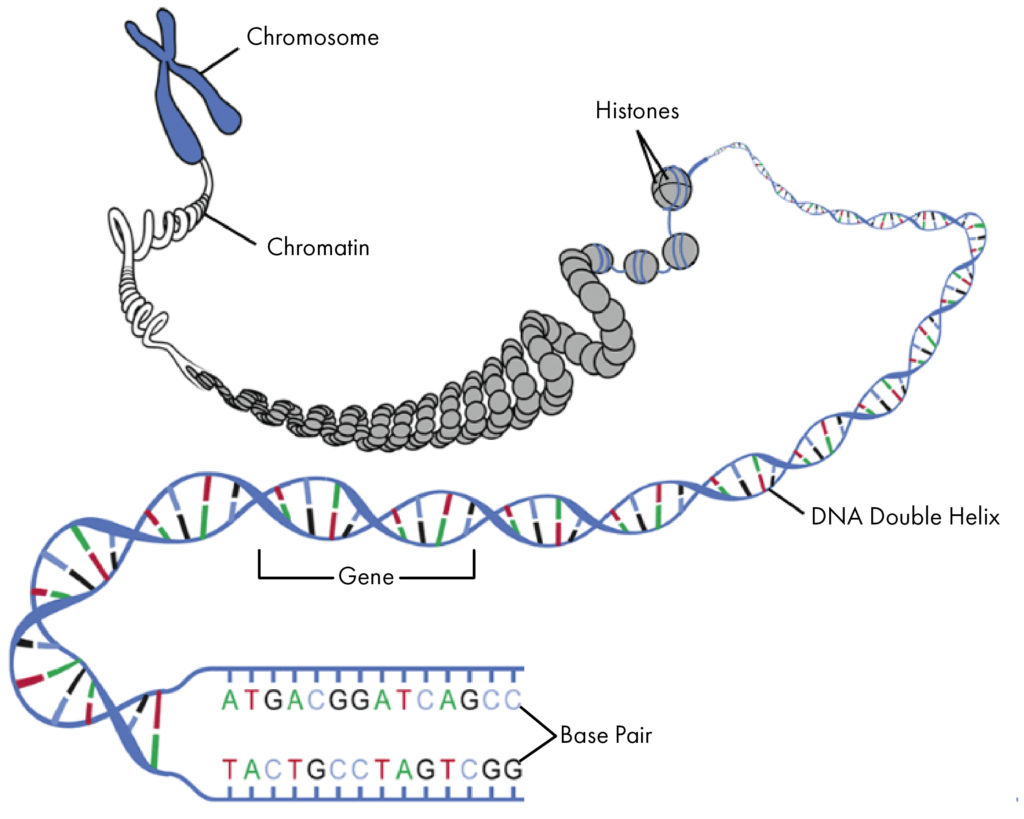
Each chromosome is composed of tightly packed chromatin; this is a structure made up of DNA (deoxyribonucleic acids), wrapped around proteins known as histones. The DNA molecule consists of two strands which wind around each other to form a double helix ‘twisted ladder’ structure.
The sides of the ladder are made up of sugars and phosphates while the ‘rungs’ of the DNA ladder are pairs of nucleic acids, known as base pairs. There are four types of bases in DNA; adenine (A), thymine (T), cytosine (C) and guanine (G). The bases pair in a specific way; A always pairs with T, and C always pairs with G.
So how does DNA provide the instructions to make us? The genetic code is formed from the bases along one strand of the DNA molecule, known as the DNA sequence. The DNA sequence is broken up into genes, units of the DNA sequence which provide the instructions for making proteins. The complete set of human DNA (3 billion base pairs!), including all genes, is known as the genome.
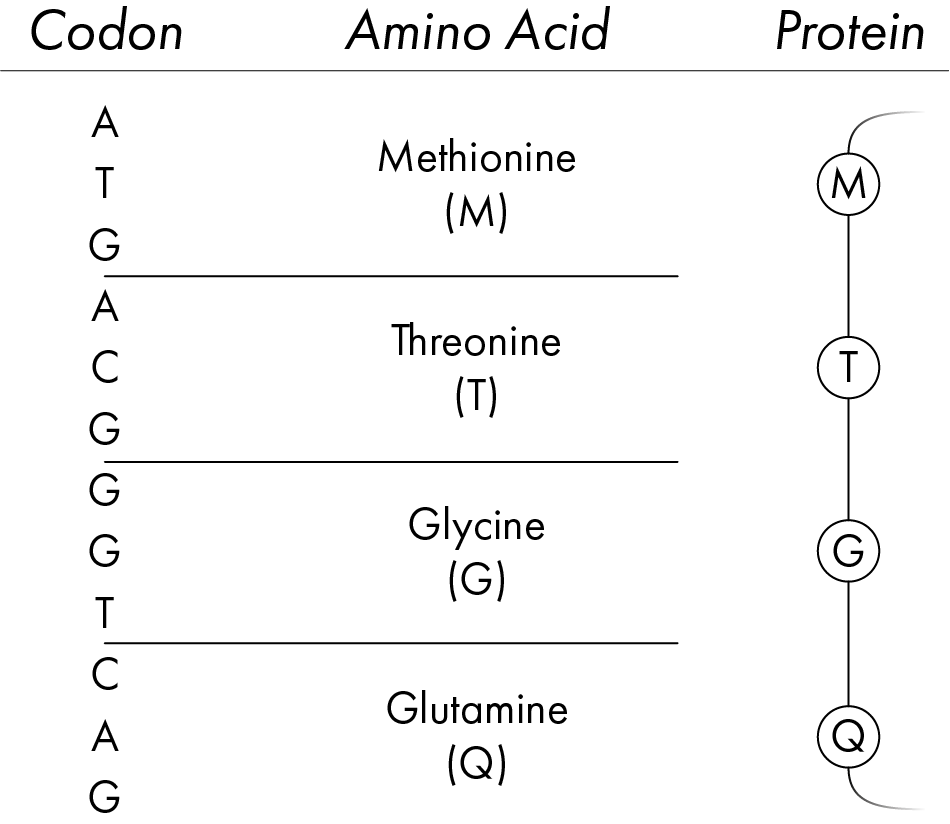
The bases A, T, C and G are the ‘letters’ of the DNA sequence, with each group of three letters forming a ‘word’ known as a codon. Each codon is associated with a specific amino acid, as the codons are ‘read’ along the DNA strand, the specific amino acid is added to a growing chain which, once complete, forms a protein. Proteins are the building blocks of life; they form the components of every one of those 30 trillion cells as well as acting in a whole host of processes that keep us alive.
Patterns of Inheritance
As we have previously discussed, we inherit half our genetic code from our mother and half from our father. Therefore, at every position in our genome we have two alleles (forms of the gene), one inherited from each parent.
The genotype is the combination of alleles we have inherited. The alleles may be the same (homozygous) or different (heterozygous) forms of a gene and can be dominant or recessive in the way the gene is expressed as a characteristic, known as the phenotype.
Autosomal Dominant
An allele is described as autosomal dominant when only one copy is required to display the phenotype associated with the gene on an autosome. The associated phenotype will be present whether the individual is heterozygous or homozygous for the allele.
Autosomal Recessive
An allele is described as autosomal recessive when two copies are required to display the phenotype associated with the gene on an autosome. The associated phenotype will be present only when the individual is homozygous for the allele. Individuals who are heterozygous for the autosomal recessive allele are described as a carrier, they will not display the phenotype but can pass the allele onto their children.
X-linked
The X chromosome contains many genes, including those not related to sex determination. While the Y chromosome does not. Therefore, genes on the X chromosome required by both sexes display a characteristic inheritance pattern described as X-linked, or sex-linked.
Females, with two X chromosomes, have two copies of each gene and can therefore be heterozygous or homozygous for an allele. Males, with one X and one Y chromosome, express all the alleles present on their only X chromosome (which they receive from their mother) and are described as hemizygous for that allele.
It’s all in the name – How variants are described
If you have delved into the world of genetic testing, knowing your bases from your amino acids is pretty important in understanding what your results mean for your genetic code, especially when reading scientific literature.
Ready to dive in?
Through a technique known as DNA sequencing, which determines the precise order of bases within the human genome, scientists now have a good idea of what the ‘common’ base is at each position, while most of the DNA sequence is the same between individuals, there are positions which are known to vary.
Most direct to consumer genetic testing is through ‘genotyping’ – this means the company providing your results has determined, at key positions across the genome, whether you have inherited an A, T, C or G base and compared this to the common base expected at this position. For example, if the common base is an A, but you are shown to have inherited a G, this is known as a variant, because it varies from the base that is most commonly found at that position. Variants are often described as single nucleotide polymorphisms (SNPs – pronounced ‘snips’). The different forms of the same gene produced by the presence of variants within the gene are termed alleles. Therefore, you will often here these terms used interchangeably, ‘I have the G variant’, ‘I have the G SNP’ or ‘I have the G allele’.
Though standard nomenclature does exist, the same variant can be described in many ways, which can be confusing.
For example, one of the variants in the CFTR gene, known to cause Cystic Fibrosis, is a change from a G base to an A base.
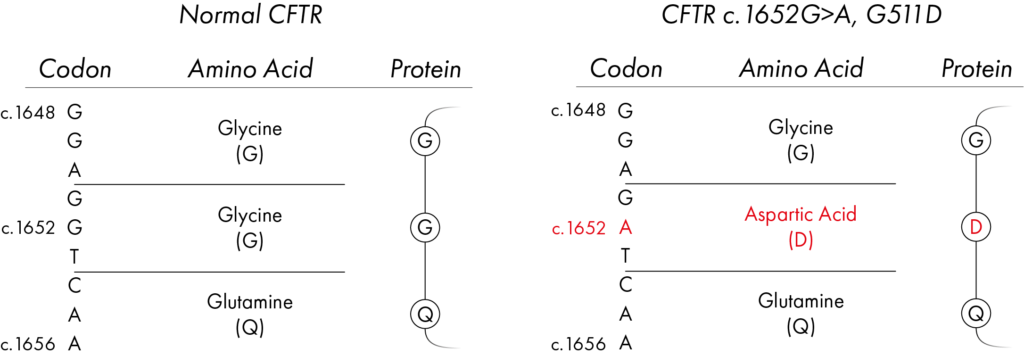
This G to A change occurs at the 1652nd position along the DNA sequence that encodes the CFTR gene. A ‘c’ is often put in front of this number to show it’s the DNA sequence e.g. c.1652.
Using the three-letter amino acid code, this is the 551st position of the amino acid chain, this three-letter codon should be ‘GGT’ specifying the amino acid Glycine but with the variant it changes to ‘GAT’, encoding the amino acid Aspartic acid. A ‘p’ is often put in front of this number to show it’s the protein coding sequence e.g. p.551.
As if this wasn’t enough to remember, amino acids can be known by their full name, a three-letter code or a one-letter code, for example; Glycine, Gly or G and Aspartic acid, Asp or D. Luckily researchers can use a table to help figure this all out:
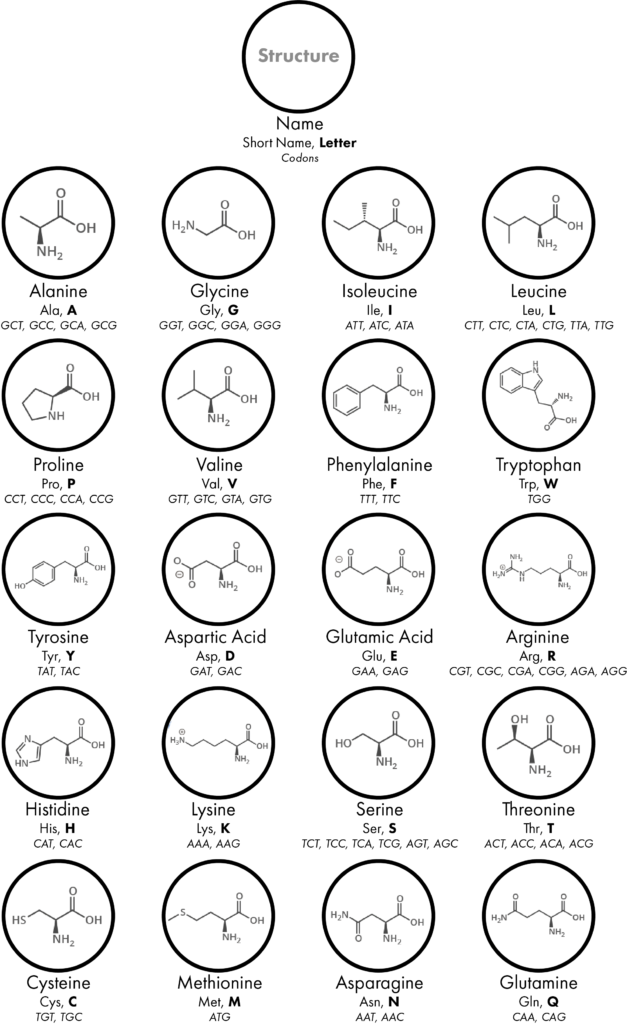
Keeping up?
Now, these numbers and letters can be arranged in a variety of ways to all represent the same thing: c.1652G>A, 1652G>A, 1652G/A, p.Gly551Asp, G551D … and a few less common ones.
Variants are also given a specific ID for data basing purposes within the scientific community, the most commonly used ID is an ‘rs number’. For example, the rs number for the CFTR gene c.1652G>A variant is rs75527207.
Variants within the genome can mean one of many things depending on the impact they have on the genetic code; they can be associated with severe diseases, give an insight into your ancestry, or indicate how you may respond to certain drugs or react to specific nutrients. Of course, the role of many variants is still not fully understood, and multiple other factors also play a large role in who you are, but at least now you should know what all these numbers and letters used to describe variants mean!
