Can Genetic Testing Show Celiac Disease?
Article at a Glance
- Genetic testing, on its own, cannot diagnose celiac disease.
- However, there are genetic markers that 95% of those with celiac disease carry, and learning you carry these genetic variants can be a strong clue that further testing is needed.
- Some variants of the HLA genes are very strongly associated with celiac disease. It is thought that in people carrying these variants, gluten is wrongly detected as a harmful pathogen. This results in the formation of auto-antibodies, leading to damage of the small intestine.

Contents
Celiac disease describes a relatively common (it affects approximately 1 in 135 of us, although complications with diagnosis may mask the true number), lifelong disorder of the digestive system, where the intestine becomes inflamed when we eat gluten.
This gives rise to the common symptoms of bloating and diarrhea, and dietary malabsorption which can cause severe complications in young children, pregnant women, and many adults.
While severe cases are often diagnosed in childhood, milder cases can exist for long periods without diagnosis, and it is possible for the disease to appear or flare up in those who previously had no symptoms.
Science doesn’t yet understand how symptoms can be so variable in different people, although there is likely to be a strong genetic element, due to a significant familial association, i.e. if your parents have it you’re likely to have it too.
Standing alone, a DNA test can’t definitively diagnose celiac disease.
However, there are genetic markers that 95% of those with celiac disease carry, but the same genes are also carried by 20-30% of the healthy population as well, so it’s important to use a multifaceted approach that combines genetic testing and traditional lab testing.
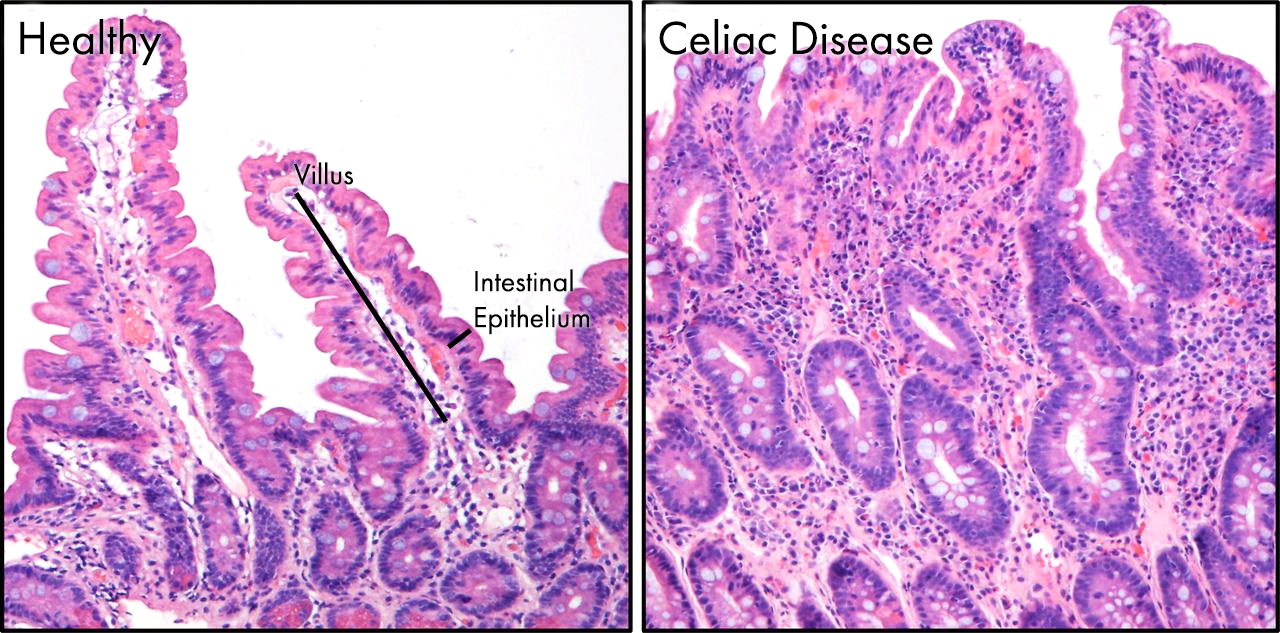
Image showing the pathology of celiac disease in the gut. On the left is a normal region of the gut epithelium. Structures called villi (singular villus) project out into the gut, massively increasing the surface area of the intestine allowing efficient absorption of nutrients. In celiac disease, these delicate villi are damaged and destroyed by the body’s immune system following its activation by gluten.
Celiac disease is an auto-immune condition
Celiac disease is an auto-immune disorder triggered by a reaction to gluten, a protein found in numerous grains such as barely and rye, with wheat in particular being the major driver.
The immune cells in the gut of people with celiac disease view gluten as a foreign pathogen. When they detect gluten in the digestive tract, they trigger the release of molecules known as auto-antibodies; specifically, anti-transglutaminase antibodies.
Antibodies are a major player in the immune system. Normally, they target foreign pathogens for destruction, however, the auto-antibodies released by celiac patients target the body’s own cells, which causes the slow and steady degradation of the villi of the small intestine.
When these cells are destroyed the individual is unable to absorb food correctly, and secretion of enzymes required to breakdown food is also impaired, thus giving rise to the common symptoms of bloating and gut problems.
For those suffering with celiac disease the only effective treatment is a strict gluten-free diet and sometimes even going gluten free isn’t enough to heal the gut.
Celiac genetic markers
If you’ve done much reading about auto-immune disorders, such as celiac disease, you’ve probably come across something known as human leukocyte antigen or HLA.
HLA is a gene-complex composed of numerous different genes which plays a key role in the modulation of our incredibly complex immune system.
When a foreign pathogen is detected, specialized immune cells engulf it and break it down into small pieces. These pieces are then presented to the surface of the immune cell by HLA proteins. This presentation then allows other immune cells to produce antibodies kicking off fully fledged immune response.
Our HLA genes are hugely variable, which is really important in making sure we’re not all susceptible to the same pathogens as each other. If we had identical HLA genes we as a species would be much more susceptible to a single potent pathogen.
So what does this have to do with celiac disease?
HLA genes and celiac disease
Simply put there are some variants of the HLA genes which are very strongly associated with celiac disease. It is thought that in people carrying these variants gluten is wrongly detected as a harmful pathogen. This results in the formation of auto-antibodies leading to damage of the small intestine.
HLA-DQ
HLA-DQ is a protein encoded for by two genes within the HLA region HLA-DQA1 and HLA-DQB1.
As discussed above, there is considerable variation in the sequence of both HLA-DQA1 and HLA-DQB1. Depending on the versions of HLA-DQA1 and HLA-DQB1 present, there are seven known serotypes, with several subtypes as well.
HLA Serotypes
Of these serotypes, and subtypes, two have been shown to have a significant association with celiac disease, DQ2.5 and DQ8, both of which are marked in the table below.[ footnote “3”] It is very important to state here that although there is a strong association (approximately 95% of celiac patients carry either the DQ2.5 or DQ8 variant), approximately 20-30% of the normal population also carry these variants. There is a third potential genetic risk marker, where DQ2.2 and DQ7 together can give rise to a disease state similar to DQ2.5.
HLA-DQ Frequencies in Celiac Disease
| Serotype | Subtype | Freq (%) |
|---|---|---|
| DQ2 | 2.5 | 13.16 |
| 2.2 | 11.08 | |
| 2.3 | 0.08 | |
| DQ4 | 4.3 | 0.15 |
| 4.4 | 2.26 | |
| DQ5 | 5.1 | 10.62 |
| 5.2 | 1.25 | |
| 5.3 | 2.03 | |
| 5.4 | 0.08 | |
| DQ6 | 6.1 | 0.66 |
| 6.2 | 14.33 | |
| 6.3 | 5.83 | |
| 6.4 | 3.40 | |
| 6.9 | 0.71 | |
| DQ7 | 7.2 | 0.05 |
| 7.3 | 6.79 | |
| 7.4 | 0.03 | |
| 7.5 | 11.06 | |
| 7.6 | 0.11 | |
| DQ8 | 8.1 | 10.55 |
| DQ9 | 9.2 | 3.66 |
| 9.3 | 0.79 |
Table showing the incredible variation in DQ serotypes and subtypes. DQ2.5 and DQ8, marked in red, are associated with celiac disease. When DQ2.2 and DQ7, shown in yellow, occur together this can equate to DQ2.5 Table adapted from.
HLA-DQ and me
Now we know that there are some genetic variants associated with celiac disease what can you do about it?
Well first, if you suspect you may have celiac disease or just want peace of mind you should contact your doctor. They offer you two non-invasive tests a blood test to look for auto-antibodies and a genetic test to check your HLA-DQ status.
As discussed above, neither of these tests are 100% effective, so if you are symptomatic for celiac disease and your tests are negative they may recommend a more invasive test such as taking a biopsy of your intestine via an endoscopy. If your results are negative, and you don’t have symptoms this is likely to be unnecessary.
23andme can’t identify celiac genes perfectly
However, I’m sure many of you have undergone some form of consumer based genetic testing like 23andme and are wondering if you can interpret your results to predict celiac risk. The short answer is not definitively. The test is not sensitive enough to accurately predict a risk, and the clinical genetic testing described above is to be preferred.
The slightly longer answer is… maybe. This study describes a means of predicting HLA-DQ serotype or subtype based on the variation of six SNPS; rs2395182, rs7775228, rs4713586, rs2187668, rs4639334 and rs7454108. There are single SNPs which can identify DQ2.5, DQ7 and DQ8. For DQ2.2, and the rare DQ4, things are a little more complex, to accurately predict for this three SNPs need to be assessed together. This can be quite complex to understand, so I’ve summarised the risk alleles in the table below.
HLA-DQ Serotype and SNPs
| Serotype | SNP ID | Risk Allele (Freq %) |
|---|---|---|
| DQ2.5 | rs2187668 | T (0.09) |
| DQ8 | rs7454108 | G (0.18) |
| DQ2.2/DQ4 See legend for more info. | rs2395182 | T (0.71) |
| rs7775228 | G (0.10) | |
| rs4713586 | A (0.95) G (0.05) |
|
| DQ7 | rs4639334 | A (0.09) |
Risk alleles of SNPs for DQ2.5, DQ7 and DQ8. DQ2.2 and DQ4 status are predicted using the same three SNPS as a haplotype, with rs4713586 being the deciding SNP. Having a T allele for rs2395182 and a G allele for rs7775228 points towards both DQ2.2 and DQ4. Then the more common A allele of rs4713586 classes as DQ2.2 and the much rarer G allele as DQ4. Table adapted from.
However, as above a proper diagnosis of celiac disease requires discussion with your doctor, and as part of that discussion a much more sensitive genetic test is likely to be recommended. So personal interpretation may be of interest, but may not be of much benefit.
Closing thoughts
I hope I’ve managed to go through celiac disease and how gluten and our own immune systems combine to give disease to the disorder.
There is a strong genetic element underpinning celiac disease, however there is not a 100% link. In other words, if you have the markers, you likely have celiac, but it’s not a certainty based on genetics.
Therefore, if you feel you may be suffering from celiac disease the best course of action is to discuss with your doctor, and ensure you get a proper panel of testing performed.



